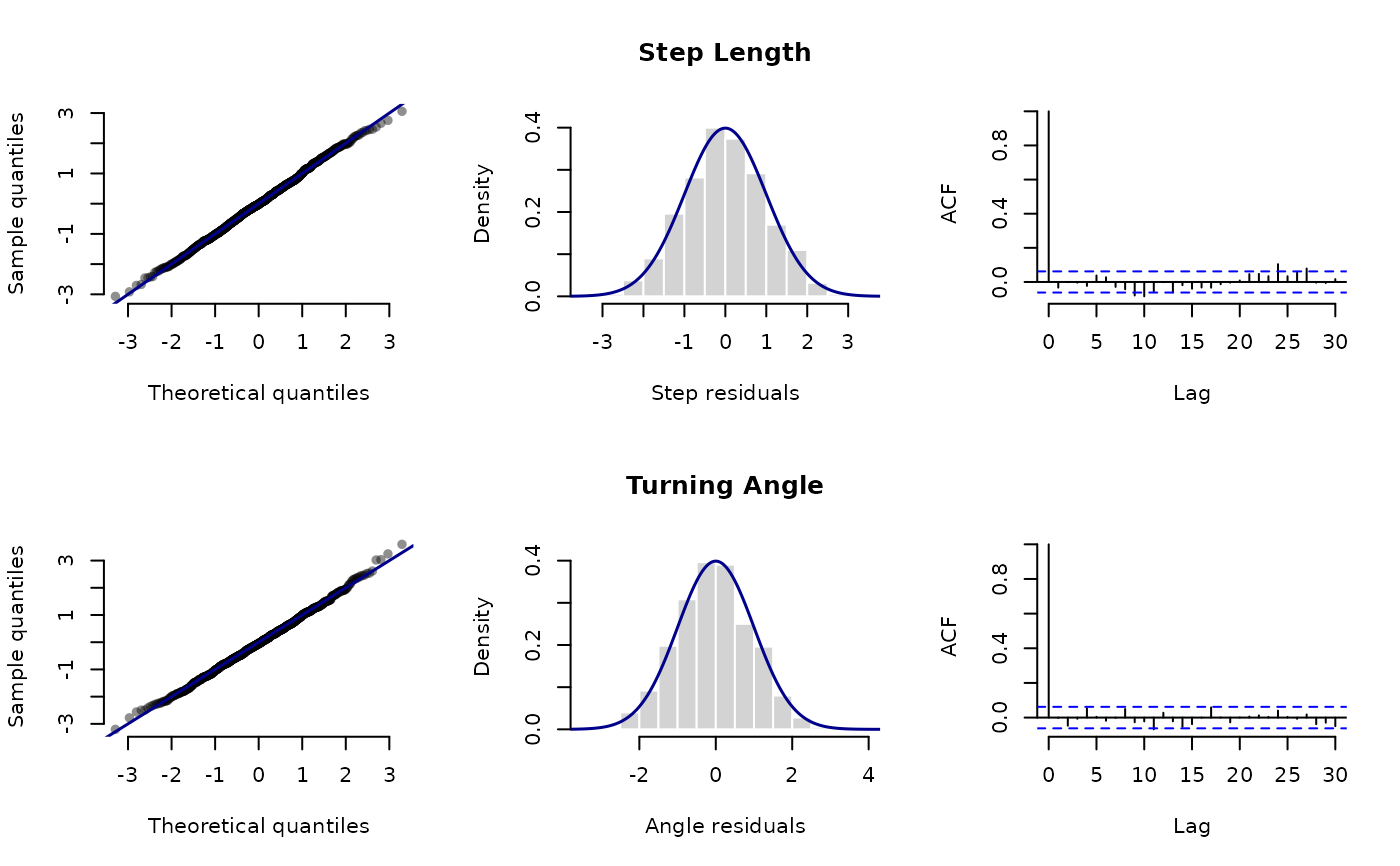
Plot pseudo-residuals
plot.LaMaResiduals.RdPlot pseudo-residuals computed by pseudo_res.
Arguments
- x
pseudo-residuals as returned by
pseudo_res- col
character, color for the QQ-line (and density curve if
histogram = TRUE)- lwd
numeric, line width for the QQ-line (and density curve if
histogram = TRUE)- main
optional character vector of main titles for the plots of length 2 (or 3 if
histogram = TRUE)- breaks
breaksargument passed to hist- axis.lab
labels used for the x and y axis of each plot (named list)
- ...
currently ignored. For method consistency
Examples
## pseudo-residuals for the trex data
step = trex$step[1:1000]
angle = trex$angle[1:1000]
nll = function(par){
getAll(par)
Gamma = tpm(logitGamma)
delta = stationary(Gamma)
mu = exp(logMu); REPORT(mu)
sigma = exp(logSigma); REPORT(sigma)
kappa = exp(logKappa); REPORT(kappa)
allprobs = matrix(1, length(step), 2)
ind = which(!is.na(step) & !is.na(angle))
for(j in 1:2) {
allprobs[ind,j] = dgamma2(step[ind], mu[j], sigma[j]) *
dvm(angle[ind], 0, kappa[j])
}
-forward(delta, Gamma, allprobs)
}
par = list(logitGamma = c(-2,-2),
logMu = log(c(0.3, 2.5)),
logSigma = log(c(0.3, 0.5)),
logKappa = log(c(0.2, 1)))
obj = MakeADFun(nll, par)
opt = nlminb(obj$par, obj$fn, obj$gr)
#> outer mgc: 1346.261
#> outer mgc: 327.3472
#> outer mgc: 200.2404
#> outer mgc: 215.4124
#> outer mgc: 65.44812
#> outer mgc: 55.5835
#> outer mgc: 89.48845
#> outer mgc: 50.01703
#> outer mgc: 19.29538
#> outer mgc: 48.04218
#> outer mgc: 12.58283
#> outer mgc: 8.252805
#> outer mgc: 12.50345
#> outer mgc: 27.17566
#> outer mgc: 21.5893
#> outer mgc: 18.1972
#> outer mgc: 26.14859
#> outer mgc: 8.841267
#> outer mgc: 13.26384
#> outer mgc: 8.94153
#> outer mgc: 3.621365
#> outer mgc: 9.520088
#> outer mgc: 5.015184
#> outer mgc: 7.817317
#> outer mgc: 15.53745
#> outer mgc: 6.502823
#> outer mgc: 9.099285
#> outer mgc: 2.16175
#> outer mgc: 8.943538
#> outer mgc: 0.765158
#> outer mgc: 3.67778
#> outer mgc: 1.449207
#> outer mgc: 3.683187
#> outer mgc: 4.210138
#> outer mgc: 2.460138
#> outer mgc: 2.917419
#> outer mgc: 1.790974
#> outer mgc: 0.8606995
#> outer mgc: 1.553886
#> outer mgc: 0.5824252
#> outer mgc: 0.2281493
#> outer mgc: 0.0414528
#> outer mgc: 0.02232808
#> outer mgc: 0.0191833
#> outer mgc: 0.02028886
#> outer mgc: 0.009637611
#> outer mgc: 0.003269403
#> outer mgc: 0.0006318679
#> outer mgc: 7.4312e-05
mod = report(obj)
pres_step = pseudo_res(step, # observations
"gamma2", # family that is used
list(mean = mod$mu, sd = mod$sigma), # the family's parameters
mod = mod) # model object
pres_angle = pseudo_res(angle,
"vm",
list(mu = 0, kappa = mod$kappa),
mod = mod)
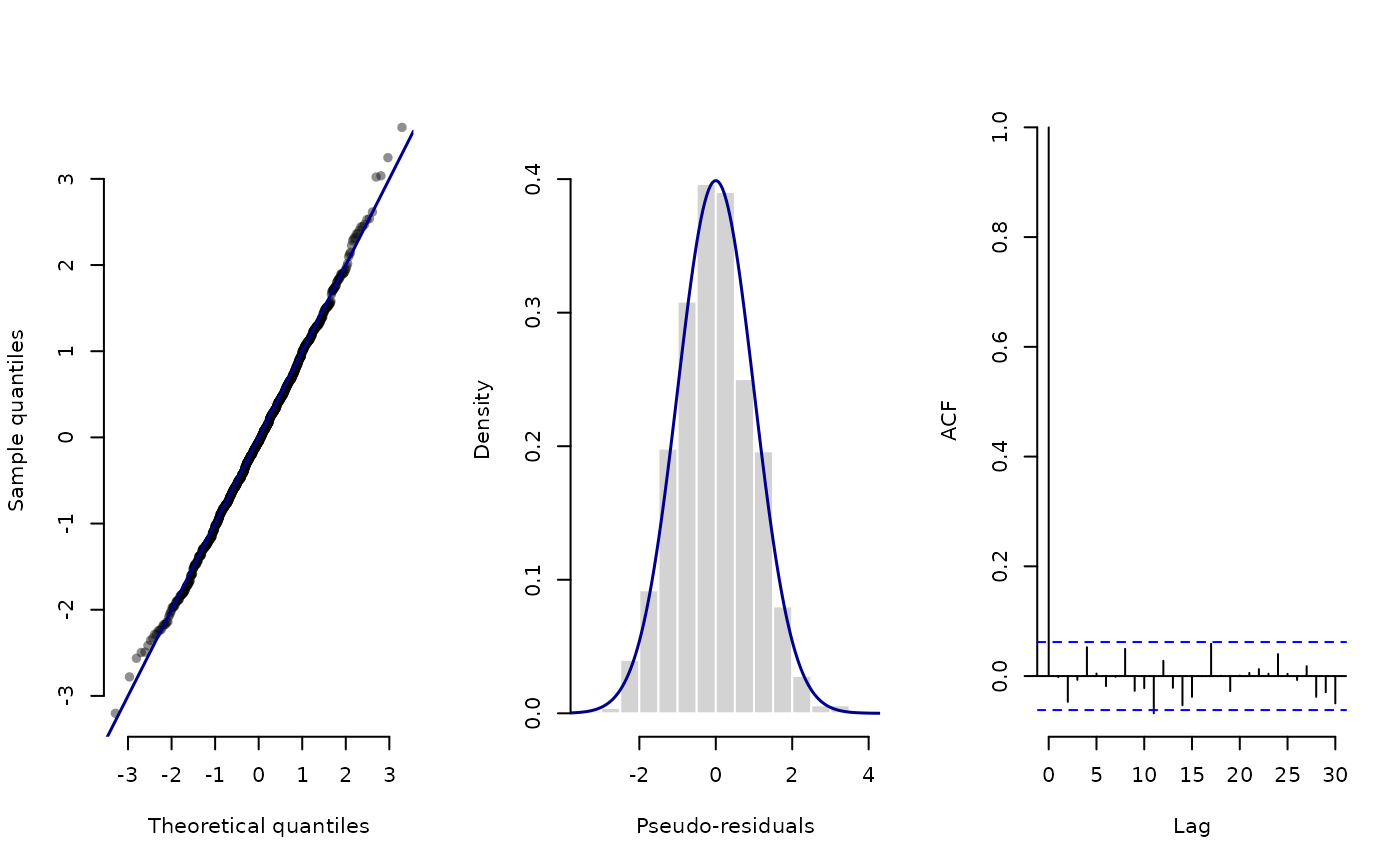
# separate plots
plot(pres_step)
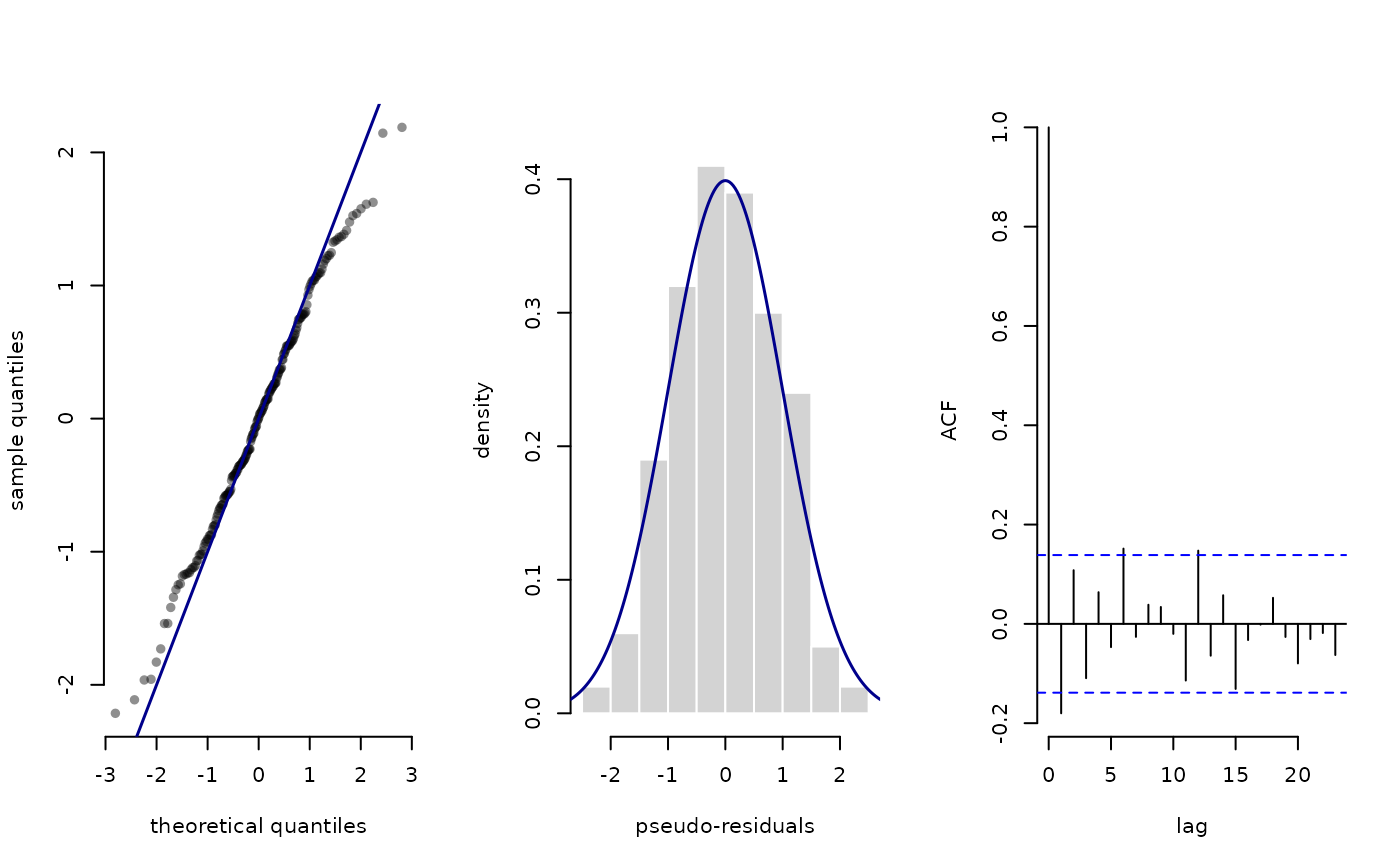 plot(pres_angle)
plot(pres_angle)
 # together
par(mfrow = c(2,3))
plot(pres_step, main = c("", "Step Length", ""),
axis.lab = list(hist = c("Step residuals", "Density")))
plot(pres_angle, main = c("", "Turning Angle", ""),
axis.lab = list(hist = c("Angle residuals", "Density")))
# together
par(mfrow = c(2,3))
plot(pres_step, main = c("", "Step Length", ""),
axis.lab = list(hist = c("Step residuals", "Density")))
plot(pres_angle, main = c("", "Turning Angle", ""),
axis.lab = list(hist = c("Angle residuals", "Density")))