Calculate pseudo-residuals
pseudo_res.RdFor HMMs, pseudo-residuals are used to assess the overall goodness-of-fit of the model. These are based on the cumulative distribution function (CDF) $$F_{X_t}(x_t) = F(x_t \mid x_1, \dots, x_{t-1})$$ and can be used to quantify whether an observation is extreme relative to its model-implied distribution.
This function calculates such residuals via the probability integral transform, based on the local state probabilities obtained by stateprobs or stateprobs_g and the respective parametric family.
Usage
pseudo_res(
obs,
dist,
par,
stateprobs = NULL,
mod = NULL,
normal = TRUE,
discrete = NULL,
randomise = TRUE
)Arguments
- obs
vector of continuous-valued observations (of length
nObs)- dist
character string specifying which parametric CDF to use (e.g.,
"norm"for normal or"pois"for Poisson) or CDF function to evaluate directly. If a discrete CDF function is passed, thediscreteargument needs to be set toTRUEbecause this cannot determined automatically.- par
named parameter list for the parametric CDF
Names need to correspond to the parameter names in the specified distribution (e.g.
list(mean = c(1,2), sd = c(1,1))for a normal distribution and 2 states). This argument is as flexible as the parametric distribution allows. For example you can have a matrix of parameters with one row for each observation and one column for each state.- stateprobs
matrix of local state probabilities for each observation (of dimension c(nObs, nStates)) as computed by
stateprobs,stateprobs_gorstateprobs_p- mod
optional model object containing required quantities
When using automatic differentiation either with
RTMB::MakeADFunorqremland includeforward,forward_gorforward_pin your likelihood function, the objects needed for state decoding are automatically reported after model fitting. Hence, you can pass the model object obtained from runningreport()or fromqremldirectly to this function and avoid calculating local state proabilities manually. In this case, a call should look likepseudo_res(obs, "norm", par, mod = mod).- normal
logical, if
TRUE, returns Gaussian pseudo residualsThese will be approximately standard normally distributed if the model is correct.
- discrete
logical, if
TRUE, computes discrete pseudo residuals (which slightly differ in their definition)By default, will be determined using
distargument, but only works for standard discrete distributions. When used with a special discrete distribution, set toTRUEmanually.- randomise
for discrete pseudo residuals only. Logical, if
TRUE, return randomised pseudo residuals. Recommended for discrete observations.
Value
vector of pseudo residuals of class LaMaResiduals or list containing lower, upper, and mean if discrete residuals are not randomised.
Details
Pseudo-residuals are based on the probability integral transform. If the cumulative distribution function (CDF) \(F\) of a random variable \(X\) is known, then \(F(X)\) is uniformly distributed on \([0,1]\). Applying the standard normal quantile function \(\Phi^{-1}\) then gives a residual that is approximately standard normal under the model.
For discrete-valued observations, the CDF is a step function, so the residual
is not uniquely defined. One can compute a "lower" and "upper" pseudo-residual
corresponding to \(F(X-1)\) and \(F(X)\), respectively.
If randomise = TRUE, a value is drawn uniformly
between the lower and upper bounds. In this case, the resulting pseudo-residuals
are again approximately standard normal after applying the standard normal quantile function.
See also
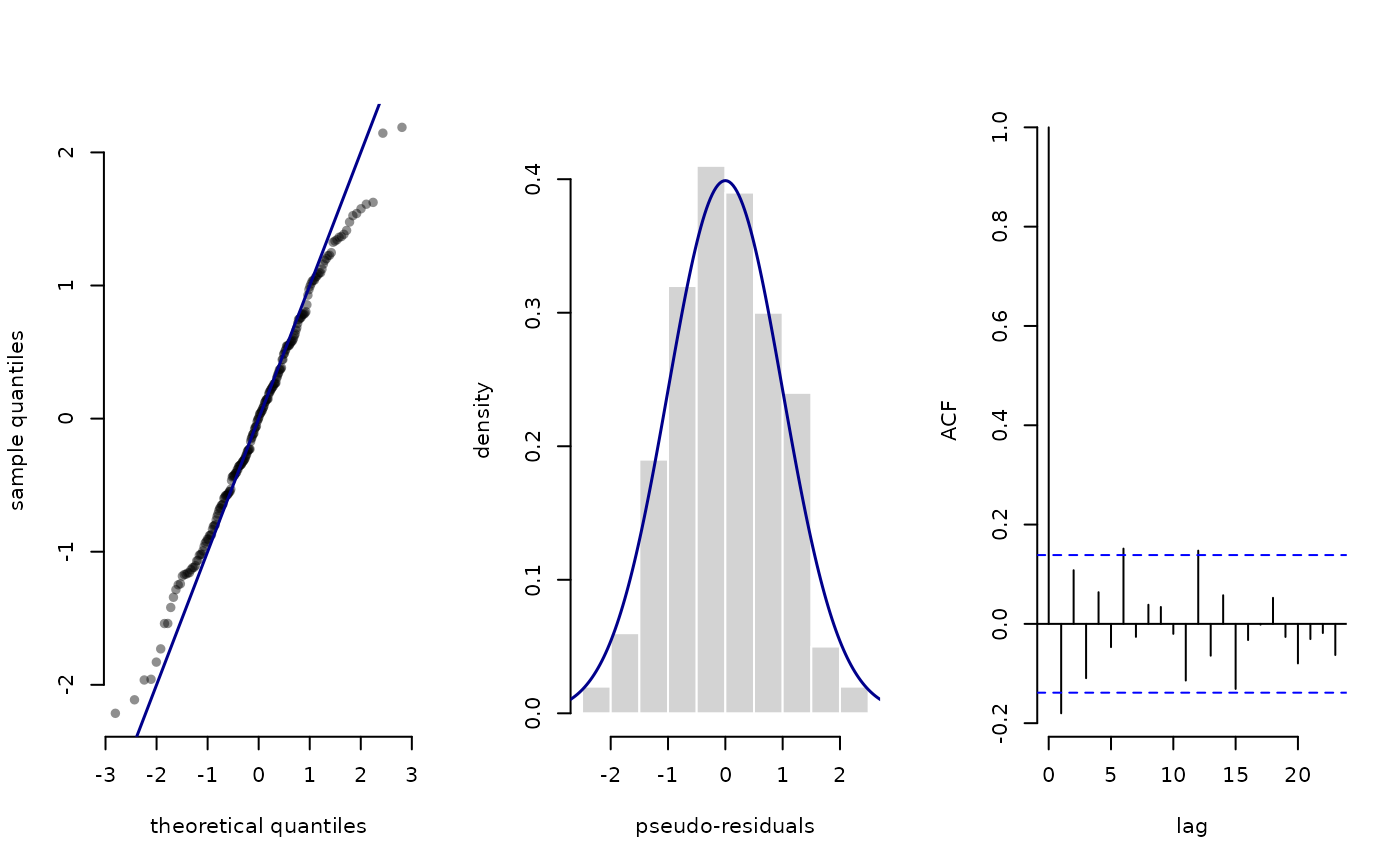
plot.LaMaResiduals for plotting pseudo-residuals.
Examples
## continuous-valued observations
obs = rnorm(100)
stateprobs = matrix(0.5, nrow = 100, ncol = 2)
par = list(mean = c(1,2), sd = c(1,1))
pres = pseudo_res(obs, "norm", par, stateprobs)
## discrete-valued observations
obs = rpois(100, lambda = 1)
par = list(lambda = c(1,2))
pres = pseudo_res(obs, "pois", par, stateprobs)
#> Calculating discrete pseudo-residuals
#> Randomised between lower and upper
## custom CDF function
obs = rnbinom(100, size = 1, prob = 0.5)
par = list(size = c(0.5, 2), prob = c(0.4, 0.6))
pres = pseudo_res(obs, pnbinom, par, stateprobs,
discrete = TRUE)
#> Calculating discrete pseudo-residuals
#> Randomised between lower and upper
# if discrete CDF function is passed, 'discrete' needs to be set to TRUE
## no CDF function available, only density (artificial example)
obs = rnorm(100)
par = list(mean = c(1,2), sd = c(1,1))
# construct CDF using numerical integration
cdf = function(x, mean, sd) integrate(dnorm, -Inf, x, mean = mean, sd = sd)$value
pres = pseudo_res(obs, cdf, par, stateprobs)
#> Assuming 'dist' evaluates a continuous CDF. Set 'discrete = TRUE' if not.
### Full model fit example ###
step = trex$step[1:200]
nll = function(par){
getAll(par)
Gamma = tpm(logitGamma)
delta = stationary(Gamma)
mu = exp(logMu); REPORT(mu)
sigma = exp(logSigma); REPORT(sigma)
allprobs = matrix(1, length(step), 2)
ind = which(!is.na(step))
for(j in 1:2) allprobs[ind,j] = dgamma2(step[ind], mu[j], sigma[j])
-forward(delta, Gamma, allprobs)
}
par = list(logitGamma = c(-2,-2),
logMu = log(c(0.3, 2.5)),
logSigma = log(c(0.3, 0.5)))
obj = MakeADFun(nll, par, silent = TRUE)
opt = nlminb(obj$par, obj$fn, obj$gr)
mod = obj$report()
pres = pseudo_res(step, # observation sequence
"gamma2", # parametric family that was used
list(mean = mod$mu, sd = mod$sigma), # parameters for that family
mod = mod) # model object
plot(pres)